Imagine if all the music on itunes was interrupted by blocks of white noise that you had to manually edit out every time you downloaded a song. Makes no sense right? Why would songs be recorded that way? Why not just remove the junk at the source, so you don’t have to waste time and energy?
That’s exactly what scientists wondered when they discovered genetic material known as introns.
What are introns?
Genes are regions of DNA that are used as templates to make molecular messages called RNA. RNA provides instructions to make the proteins that determine an organism’s traits. This process, called “gene expression” is the “central dogma” of molecular biology. Before RNA is used to make proteins, large sections called introns are removed. The remaining sections, which actually contain the genetic information, are spliced back together.
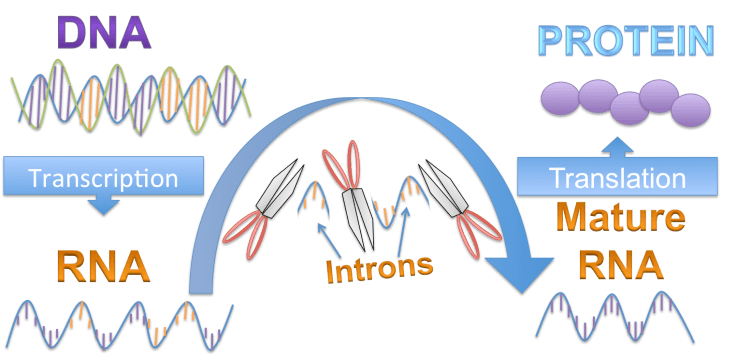
Introns, originally called “junk DNA“, are not present in the mature RNA and do not influence the final protein product. So why do cells waste energy, copying introns just to remove them immediately afterwards?
The function of this “junk DNA”
Returning to the music analogy, imagine itunes gives in to complaints and removes the white noise. Customers are surprised to find that suddenly some songs are not downloading as efficiently. Some can only be downloaded a few times per day, others will not download at all, and the most popular songs are hit the hardest
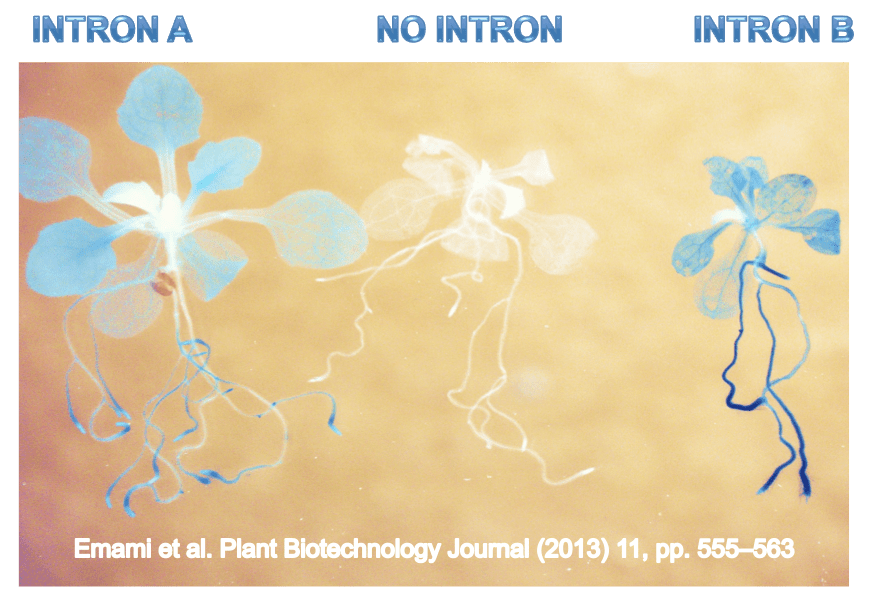
That’s what happens when scientists try to express genes without introns. For many genes, the intronless version does not yield as many copies of RNA. Further, some genes are completely dependent on this “intron-mediated enhancement” or “IME”. Only certain introns exhibit IME, and they tend to be found in the most fundamental genes, those that are expressed all the time.
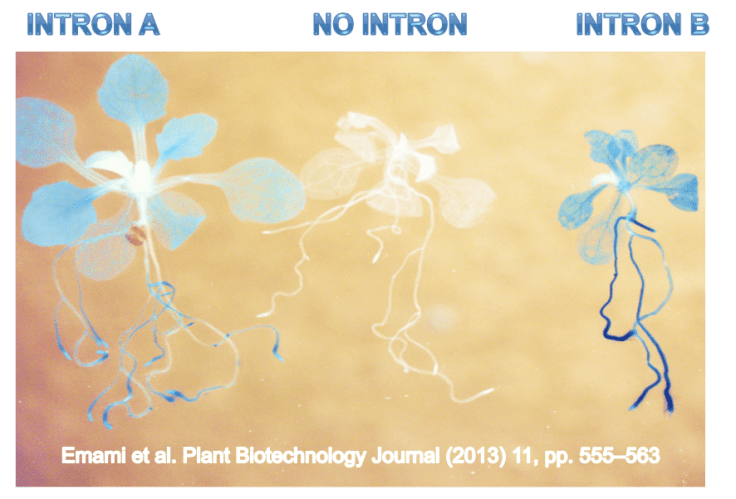
Introns can play a hugely important role
IME can increase the expression of genes anywhere from 2 to 20 fold in plants, animals, and fungi. In my lab, we study IME in plants, because introns are routinely used to increase expression of genes in crops from herbicide-tolerant soybeans to nutritionally enhanced rice. Better understanding IME could enable us to design synthetic introns that maximize gene expression potential.
Despite the obvious utility of IME, little is known about the mechanism. What we do know doesn’t fit conventional models of gene expression. For example, in order to exhibit IME, the intron must be very close to the start of the gene. Remarkably, if an IME intron is inserted into a gene that is usually only expressed in certain tissue, it can cause that gene to be expressed everywhere. New data shows that IME can even influence the length of an RNA message and act independent of DNA regions normally thought to be essential for expression.
If you want to dig deeper, my advisor Dr. Alan Rose and I published a review on the topic aptly titled “the enduring mystery of intron-mediated enhancement”.
